XCP Imaging Pipeline¶
Warning
xcpEngine only support fmriprep outputs with MNI152NLin2009cAsym template
The XCP imaging pipeline (XCP system) is a free, open-source software package for processing of multimodal neuroimages. The XCP system uses a modular design to deploy analytic routines from leading MRI analysis platforms, including FSL, AFNI, and ANTs.
The XCP system is designed to run in the Linux bash shell or from a Docker or Singularity image. We
strongly recommend using Docker or Singularity. Users provide XCP Engine with the output from
FMRIPREP and specify parameters for the analysis that they wish to perform. XCP Engine parses
the user-provided parameters to build and run a processing pipeline. The XCP system supports a
number of pipeline modalities, including functional connectivity, structural processing, and task fMRI.
Useful Features¶
XCP Engine provides tools to take FMRIPREP output and perform the next steps required for many
functional connectivity and structural analyses. FMRIPREP is required for initial preprocessing
of fMRI data; XCP performs denoising and computes additional derivatives used for analyses. For
structural processing, either FMRIPREP or the dedicated struct module in XCP can be used.
Neuroimage processing¶
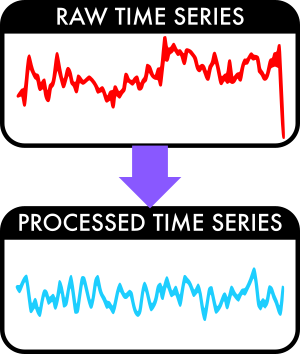
Neuroimage processing refers collectively to the set of strategies used to convert the “raw” images collected from the scanner into appropriate inputs to group-level statistical analyses. The results of any group-level neuroimaging analysis will be sensitive to the combination of strategies used to process the subject-level images. Reproducibility of processing strategies is therefore critical.
MRI time series collected from the scanner are typically noisy. Processing mitigates the effect of artefact and moves all subject-specific images into a common atlas space to facilitate direct voxelwise comparison.
It encompasses denoising, filtering, registration, and production of any subject-level derivative maps (for instance, seed-based connectivity maps or task contrasts). Processing includes production of summary measures on both a voxelwise and ROI-wise basis. Notably, processing in XCP does not include group-level statistical analysis.
Processing pipelines¶
The XCP system was designed with the importance of scientific reproducibility in mind. Often, research groups process neuroimages using a dedicated, standalone “pipeline” for each image modality. For instance, task activation, perfusion, and functional connectivity analyses might each be processed by a separate script.
However, this approach can easily lead to inconsistencies in output directory conventions, difficulty tracking pipeline versions, and limited flexibility in updating pipelines, all of which ultimately combine to compound reproducibility challenges. Often, many common routines are deployed across multiple MRI modalities, and it is in the interest of the investigator to minimize redundancy and maximize reproducibility.
The modular, atomic design of the XCP system allows for streamlined reproduction and recombination of frequently used image processing strategies across multiple pipelines and modalities.
Features¶
The XCP system aims to provide a multimodal library of common processing routines that will improve scientific reproducibility. Features include:
- Compatability with
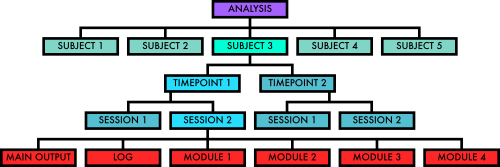
FMRIPREP- Standardized output directory structure and naming conventions
- Systematized quality control variables computed and collated for each analysis (for easy identification of motion and registration outliers)
- Region-wise quantification of any voxelwise derivative map for any number of parcellation schemes or regions of interest
- Easy addition of new regions of interest, node systems, or parcellation schemes
- Analyses either in standard/atlas or subject/native space
Contents¶
- Overview
- Pipeline configuration
- xcpEngine containers (Extra Info)
- Modules
- Utilities
- Developing/Debugging XCP
- Quality Control Dictionary
- xcpEngine on Flywheel